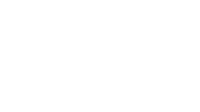Marc Karnat
Deep-Learning Epithelial Tissue Mechanics (DELET’ME) project
Team: Out-of-equilibrium Mechanics (OM) team (CPT - CENTURI)
His background
December 2020 - present | CENTURI PhD student
2019 - 2020 | Cell Physics Master II - University of Strasbourg (France)
2017 - 2020 | Engineering degree in Physics and modelling department, specializing in cell physics - Telecom Physique Strasbourg (France)
2015 - 2017 | Preparatory classes specialized in Physics and Science of Engineering - Chrestien de Troyes (France)
Contact
About his PhD project
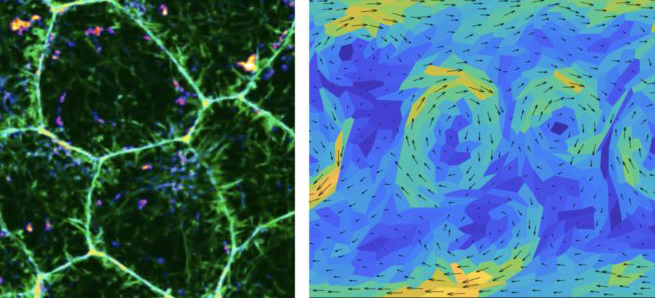
A large proportion of cancers come from epithelial tissues whose function is to protect inner organs. Our objective is to build an algorithm to infer the mechanical properties of these epithelial tissues based on optical images. As the rigidity of a tissue can be used as a clinical criterion for the recognition of pre-cancerous tissues, such algorithm could provide a new clinical insight to live endoscopies. At the OM theory group, we aim at understanding the physics of living materials based on analytical (e.g. active liquid crystal theories) and numerical models, e.g. vertex model, whereby the overall tissue dynamics results from the mechanical balance between a set of representative cellular forces – namely, viscous resistance, interfacial tension and cellular pressure (see image below). We put a strong emphasis on understanding the role of out-of-equilibrium fluctuations in tissue dynamics. These are now increasingly recognized as playing a crucial role in controlling whether a material is rigid and elastic,
or soft and fluid; yet, so far, the contribution of active force fluctuations has been significantly overlooked in tissue mechanics models. To fill this gap, we propose new out-of-equilibrium statistical physics tools that are predictive, shedding new lights on the role that local biochemical perturbations can play in the global mechanical tissue properties. During this PhD, we propose to use our fluctuating vertex model - with well-controlled mechanical properties - to expand the capability of a deep learning code currently developed in the team with the objective to infer mechanical properties of our simulated tissues.